AEROBIC BIODEGRADATION OF XYLENE WITH BIOMASS INCREASE
Biodegradation relies on the catalyzing action of enzymes produced by microbes. Therefore,
the maximal rate depends on the concentration of the enzymes or the bacteria. The variable
kmax in the Monod rate equation (see the
phenol example) can be rewritten to
account explicitly for the concentration of bacteria:
 where μmax is the specific bacterial growth rate
(s-1), B is biomass (mol C/L) and Y is the yield factor
(mol biomass-C / mol substrate-C) which expresses how much of substrate is converted into biomass.
where μmax is the specific bacterial growth rate
(s-1), B is biomass (mol C/L) and Y is the yield factor
(mol biomass-C / mol substrate-C) which expresses how much of substrate is converted into biomass.
As substrate is degraded and transformed into energy and organic molecules which can be used for
biosynthesis, the number of bacteria increase:
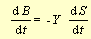 The 2 equations are coupled. Biodegradation augments when the bacteria increase in number, and
more biodegradation produces more bacteria. The decrease of substrate concentration first becomes
notable when sufficient microbes are present, and the process stops when all the substrate
has been consumed.
The 2 equations are coupled. Biodegradation augments when the bacteria increase in number, and
more biodegradation produces more bacteria. The decrease of substrate concentration first becomes
notable when sufficient microbes are present, and the process stops when all the substrate
has been consumed.
Schirmer et al. (1999, JCH 37, 69) measured aerobic breakdown of xylene in a batch experiment with pristine aquifer
sediment in which the microbes had to adapt and grow before they could digest their new food.
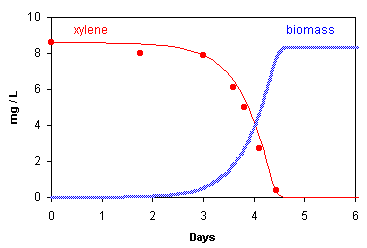
The PHREEQC input file given in Example 10.8
calculates the lines in the graph, the datapoints are from Schirmer et al. Quite conspicuous is the delay of xylene breakdown
that occurs because the microbes which degrade the substance first must grow and increase in number. When the biomass
is sufficient, xylene is quickly gone. Note that the xylene concentration in Schirmer's experiment
indicates only part of total xylene. Other biodegradable forms comprise xylene that is sorbed to the sediment and gaseous
in the headspace. These are mimicked with a retardation factor R, but they can be included as process-dependent
entities in the model. The yield factor was found to be 0.305 mol biomass-C/xylene-C. Thus, the biomass increase is:
(8.6 mg xylene/L) / (106 g/mol xylene) x (8 mol C/mol xylene)
x (R = 1.86) x (Y = 0.305)
x
(22.6 g/mol CH1.4O0.4N0.2)
= 8.32 mg biomass/L,
where CH1.4O0.4N0.2 is
the formula used for calculating the molar weight of dry biomass.
For methanogenic conditions, Y is small, about 0.05. It can be calculated that the anaerobic degradation of 40 mg
phenol produces 2.9 mg biomass with this yield factor, which is about the increase shown in the phenol experiment. Accordingly, the biomass can be considered constant in the phenol model, and
was included in the value of kmax.
